cookbook
From autoplot.org
The Cookbook is a set of user-submitted entries on how to do handy operations in Autoplot.
1. Address bar
1.1. vap file modifiers
A vap file may be modified in the address bar by adding ? and the node name:
http://autoplot.org/data/vap/PO_H0_HYD0.vap?timerange=1999-05-30
since this vap shows the time range editor instead of the dataset selector, we change multiple nodes:
http://autoplot.org/data/vap/PO_H0_HYD0.vap?options.useTimeRangeEditor=false&timerange=1999-05-30
2. Complex Configurations
The folks at LANL asked for a page with images associated with complex plot panel configurations or styles and the procedure to create them: Cooking at LANL. See also http://autoplot.org/developer.renderTypes
2.1. L-shell as Y axis
This script shows how the data is reformed to make a spectrogram with Time and LShell (sometimes called LvsT plot). This script shows how data is read in from two sources and combined.
reset() tr= getParam( 'timerange', '2014-03', 'parameter timerange' ) monitor.setTaskSize(100) monitor.started() fdpu= getDataSet('https://rbsp-ect.newmexicoconsortium.org/data_pub/rbspa/rept/level3/pitchangle/$Y/rbspa_rel03_ect-rept-sci-L3_$Y$m$d_v$(v,sep).cdf?FPDU', tr, monitor.getSubtaskMonitor(0,50,'FPDU') ) l= getDataSet('https://rbsp-ect.newmexicoconsortium.org/data_pub/rbspa/rept/level3/pitchangle/$Y/rbspa_rel03_ect-rept-sci-L3_$Y$m$d_v$(v,sep).cdf?L', tr, monitor.getSubtaskMonitor(50,70,'L Shells') ) l= synchronize( fdpu,l ) fdpu= collapse1( fdpu ) # collapse all pitch angles fdpu= fdpu[:,4] # slice at one energy lgrid= linspace( 1, 7, 40 ) monitor.setTaskProgress(80) from org.virbo.dsutil import LSpec lspec= LSpec.rebin( l, fdpu, lgrid, 1 ) ttag= lspec.property( QDataSet.DEPEND_0 ) ttag= collapse1(ttag) # collapse1 takes the average of the min and max time tags. lspec= link( ttag, lspec.property( QDataSet.DEPEND_1 ), lspec ) sliceAtL= lspec[:,10] plot(lspec,renderType='nnSpectrogram') plot(1,sliceAtL)
2.2. Events Bar under all plots
- plot the top plot http://jfaden.net/~jbf/1wire/dot4_data/$Y/$m/$d/10.592044000800.$Y$m$d.d2s?timerange=2013-01-26
- File->Add Plot, plot below http://jfaden.net/~jbf/1wire/dot4_data/$Y/$m/$d/10.952040000800.$Y$m$d.d2s?timerange=2013-01-26
- layout tab, select both plots.
- right-click, canvas->add hidden plot..
- deselect all but the x axis.
- select the new plot element, make it active
- in the address bar, "vap+inline:createEvent('2013-01-26T016:00/PT15M',0xffa0a0a0,'event1')" to make an opaque bar
Now that leaves the hidden plot above others. We want this to be below the others.
Here is a tutorial PDF of how to add events bars over two plots: http://autoplot.org/data/tutorials/20180129_events/20180129.pdf
2.3. Use Events List to control time ranges
Suppose you want to look at burst data which is collected for a few seconds every few minutes. To explore the data using the scan next and scan previous can be a bit tedious. You can use an Events List to control the time ranges. If you have a list of times formatted like so: timeStart, timeEnd, and then other columns, like the file here: http://emfisis.physics.uiowa.edu/events/rbsp-b/burst/rbsp-b_burst_times_20120910.txt. You can look at the intervals with the URI: http://emfisis.physics.uiowa.edu/events/rbsp-b/burst/rbsp-b_burst_times_20120910.txt or http://emfisis.physics.uiowa.edu/events/rbsp-b/burst/rbsp-b_burst_times_20120910.txt. Autoplot detects this special text file format with its first two time columns, and treats it is an events list. To use the data as an events list, use [menubar]->Tools->"Events List", and then enter http://emfisis.physics.uiowa.edu/events/rbsp-b/burst/rbsp-b_burst_times_20120910.txt into the address bar there. This can be made into an aggregation, and then entire set can be browsed.
TODO: There's an odd bug that the aggregation requires the "eventsListColumn" switch to be used, as in http://emfisis.physics.uiowa.edu/events/rbsp-b/burst/rbsp-b_burst_times_$Y$m$d.txt?eventListColumn=field3&column=field0&timerange=2013-03-03
3. User Interface
3.1. collection of scripts
See http://autoplot.org/data/script. Scripts that make the application do things (.jy): https://sourceforge.net/p/autoplot/code/HEAD/tree/autoplot/trunk/Autoplot/src/scripts/ Scripts that load data (.jyds): https://sourceforge.net/p/autoplot/code/HEAD/tree/autoplot/trunk/JythonDataSource/src/
3.2. editing entries from address bar recent history
- click on down arrow to show droplist of recent entries.
- use keyboard arrows to select entry to edit.
- hit escape
- edit line
- hit return or the green play button.
3.3. where's my data?
If you were looking at data recently, but can't remember where it was, you can look at Autoplot's history. Every URI plotted is recorded in $HOME/autoplot_data/bookmarks/history.txt. From the menu, you can enter a GUI to browse this file, use File->Open Recent. (Note there's no mechanism to remove old entries now. Just remove the file and restart Autoplot.) Also, adding "nohistory=true" to any URI will make it so it is not logged, and logging is not done in headless mode.
4. Labels
For a complete description of labels, see help#Axes.
4.1. Hide Axis
Right-click on axis->Axis Properties->Visible or axis->Axis Properties->"Tick Labels Visible"
4.2. IDL strings
IDL's formatting strings are supported, based on the specification of Grandle and Nystrom. With "granny" strings you can set the title to "E=mc!U2!n" or m!A2!Ns!A-2!N formats m2s-2
- !A superscript
- !B subscript
- !E superscript with smaller font size
- !D subscript with smaller font size
- !N normal position
4.3. Greek and Math Symbols
You can use greek symbols such as ρ and π in axis labels as well as unicode characters. Here are a few:
- Greek letters: Α Β Δ α β δ π ρ ω
- Math symbols: ∑ (∑) ± (±)
Here's a tool for looking up entities: http://entity-lookup.leftlogic.com/
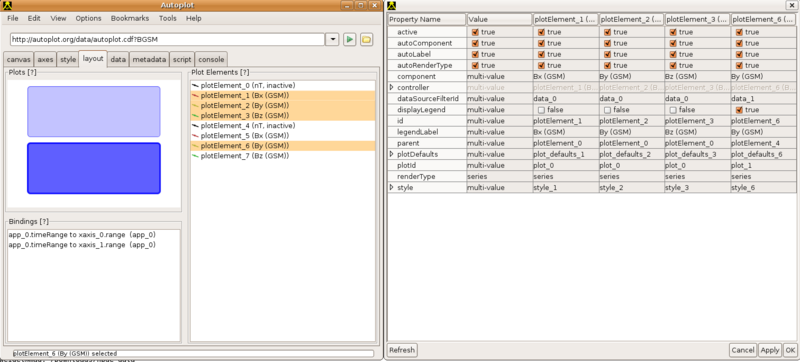
4.4. Modify Many Labels
Layout -> Plot Elements, highlight multiple elements, Right-click->Edit Plot Element Properties and a window will appear. The Value column is used to apply a setting to all other columns to its right.
4.5. Add Axis Annotations
You can add annotations to the X-axis, for labelling ephemeris. Right-click on the X-axis, select "Add additional ticks from..." and put in the name of another dataset with a common X-Axis. Autoplot will use this dataset, picking nearest neighbors to label the axis.
Note only one URI can be specified.
4.6. Background/Foreground images
The canvas object has methods for adding "decorators" which paint under all plots and above all plots. A Painter is a class with one method, paint, which takes a Java 2D graphics context calls methods for painting on it. These methods include drawImage, drawString, as well as drawing and filling shapes. This Jython script shows how images can be added to the canvas.
# Das2's canvas object has two hooks for decorating the plot, to plot below and # above the normal graphics. These provide direct access to the Graphics object # that paints the canvas. This demos how they are used. reset() import javax,java import org.das2.graph ur= 'http://cottagesystems.com/honeycomb.1.gif' image= javax.imageio.ImageIO.read( getFile( ur, monitor ) ) class BottomPaint(org.das2.graph.Painter): def paint( self, g ): g.drawImage( image, java.awt.geom.AffineTransform(), None ) class TopPaint(org.das2.graph.Painter): def paint( self, g ): g.rotate( -15*PI/180,300,300 ) g.setFont( java.awt.Font.decode( 'sans-42' ) ) g.setColor( Color( 255,255,255,80 ) ) g.drawString( "Provisional", 252,302 ) g.setColor( Color( 0,0,0,80 ) ) g.drawString( "Provisional", 250,300 ) bottom= BottomPaint() top= TopPaint() dom.canvases[0].controller.dasCanvas.removeBottomDecorators() dom.canvases[0].controller.dasCanvas.removeTopDecorators() dom.canvases[0].controller.dasCanvas.addBottomDecorator( bottom ) dom.canvases[0].controller.dasCanvas.addTopDecorator( top ) dom.plots[0].controller.dasPlot.setOpaque(True) dom.dataSourceFilters[0].uri='http://cdaweb.gsfc.nasa.gov/istp_public/data/polar/hydra/hyd_h0/$Y/po_h0_hyd_$Y$m$d_v01.cdf?ELECTRON_DIFFERENTIAL_ENERGY_FLUX&timerange=20000109'
5. Add Annotations
For several years, people have been able to add annotations to their plots. These are arbitrary text or images which are drawn on top of other components, or on top of the data within a plot. To add an annotation, right-click on a plot and select "Add Annotation", which presents some options for the annotation. Once added, the annotation can be moved and made to point at data coordinates. These annotations are then saved into .vap files as part of the canvas. Annotations can also be images, where a URL is used to locate a .gif, .jpg, or .png file.
LaTeX, a system for typesetting math equations (pronounced La-Teck), can be used with this, by first using one of many sites which render LaTeX to an image, such as quicklatex.com.
Here is a tutorial that shows how to use annotations and to draw LaTeX: HTML or as a PDF.
6. Interaction
6.1. tweaking plot position
Holding shift while mousing over a plot brings up control points that allow the plot position to be tweaked. By default Autoplot automatically adjusts the outside bounds to make room for colorbars, but this can be used to adjust heights.
6.2. time axis scan buttons
The X axis has hidden scan buttons at the beginning and end of the axis (lower right and left corners of the plot box). Mousing over these locations reveals the buttons. Note these are enabled after zooming in.
6.3. zoom with mousewheel
You can use the mouse wheel to zoom in and out. But did you know that: mouse wheel on the ends of the axis will adjust that axis in one direction. For example, put the mouse on the end of the time axis. Mouse wheeling will adjust the beginning of the axis range... Also, pressing control while spinning the wheel will adjust the range to pan along a dimension without changing the scale.
6.4. no mousewheel
The das2 graphics library was developed on Suns that didn't have mousewheels, and its controls are still around and handy when you are on a laptop without a mouse. Left click and drag along an axis to zoom in. Click and drag down slightly to zoom out. An arrow will be drawn to indicate the pending action. If you do have a middle mouse button, it will pan the axis when dragged.
7. Loading Data
7.1. watch an ASCII file
Autoplot can watch a file and get updates by polling it regularly. The following script computes the time for a wget command to execute.
#!/bin/bash SERVER="http://autoplot.org/" LOG="/tmp/webtest.log" COUNTER=0 rm -f $LOG while [ $COUNTER -lt 100000 ]; do let COUNTER=$COUNTER+1 START=`date +'%Y %m %d %H %M %S.%N'` start=$(date '+%s.%N') wget -q -O /dev/null $SERVER end=$(date '+%s.%N') delta=$(echo $end-$start | bc) echo "$START $delta" >> $LOG echo "$START $delta" sleep 1 done
It writes to the file "/tmp/webtest.log". To continuously view this file, use the undocumented (and not fully tested) feature by appending
&filePollUpdates=1&tail=100
to the URI, which will plot the last 100 lines of a file every second:
vap+dat:file:/tmp/webtest.log?time=field0&column=field6&timeFormat=$Y+$m+$d+$H+$M+$S&filePollUpdates=1&tail=100
The "&filePollUpdates=1" will work with almost any file-based data reader.
7.2. handling time ranges with implicit fields
You can have implicit fields, so that the time parser can work in more cases. Suppose you have a file 2000-03-04.dat which contains for the first field $H-$M-$S. Here's a few lines of the file http://autoplot.org/data/noDate.dat:
09:45:23 3.4 09:46:26 4.5 09:47:22 5.6
You could use implicit fields to parse this:
http://autoplot.org/data/noDate.dat?column=field1&timeFormat=$(H;Y=2000;m=3;d=4):$M:$S&time=field0
7.3. Load in only data within an interval (not "at least")
By default, Autoplot will load at least the data within the interval specified, which can result in surprises and must be dealt with in some analysis. For example, the following script will load a day's worth of data, even though only three hours are requested:
tr = '2017-11-28T12:00 to 15:00' l_erg= getDataSet('vap+cdaweb:ds=ERG_ORB_L2&filter=erg&id=pos_eq[:,0]',tr) plot(l_erg)
To fix this, a "trim" command must be inserted:
tr = '2017-11-28T12:00 to 15:00' l_erg= getDataSet('vap+cdaweb:ds=ERG_ORB_L2&filter=erg&id=pos_eq[:,0]',tr) l_erg= trim( l_erg, tr ) plot(l_erg)
8. Launching (OS-Specific)
8.1. Associate .vap files in Gnome
- right-click on a file (.vap or .cdf, for example.)
- Open with... Other application...
- Use custom command...
- Enter "/usr/local/java/bin/javaws http://autoplot.org/autoplot.jnlp -open ". (Look on your system for Java.)
- Autoplot should now be the default mechanism to open this file. The association is described in ~/.local/share/applications.
8.2. Using MIME types to launch
At the Radio and Plasma Wave Group, we defined vap to be MIME type application/x-autoplot-vap+xml so this could be associated with the Autoplot application. We have a single-jar version of autoplot, but Webstart should work as well, with something like: /usr/local/jdk8/bin/javaws http://autoplot.org/autoplot.jnlp -open %S (I think we might have to make a unix script to wrap this...)
9. Scripting
See also scripting, developer.scripting, http://autoplot.org/data/jyds, and http://autoplot.org/data/tools/.
See script.cookbook which is just for scripts.
Last, there's a github repository with hundreds of searchable examples, see https://github.com/autoplot/dev/
9.1. Animation
This loops over the day 2012-11-01, shifting to 2012-11-02, making pngs at one minute intervals:
# plot vap or uri to configure the display plot( 'vap+das2server:http://emfisis.physics.uiowa.edu/das/das2Server?dataset=rbsp/RBSP-A/HFR_spectra.dsdf&start_time=2012-11-01T00:00:00.000Z&end_time=2012-11-02T00:00:00.000Z' ) tr= DatumRangeUtil.parseTimeRange('2012-11-01') onemin= Units.seconds.createDatum(60) count= 24*60 monitor.started() i=0 monitor.setTaskSize(count) while ( i<count ): dom.plots[0].xaxis.range= tr writeToPng( '/tmp/anim20121110/%5.5d.png' % i ) i=i+1 tr= DatumRange( tr.min().add(onemin), tr.max().add(onemin) ) monitor.setTaskProgress(i) monitor.finished()
For creating a movie, see #Add_to_Jython_Search_Path.
9.2. Browse a timeseries
Any jyds script with getParam('timerange') has the "Time Series Browse" capability, and it will be added when the jython data source is loaded. See examples in http://autoplot.org/data/jyds.
9.3. getParam
The function getParam(param,default,label) is a special function that allows arguments to be passed into a script. When a script is edited and getParam is found, and GUI is automatically created. getParam is used in all contexts: for data sources, it gets named arguments like &p=1&s=a. For command-line scripts (--script=), command line parameters are passed in. For application scripts, the default value is used unless ALT (option on a Mac) is held down with the Execute button.
9.4. Take the average of many images
David at Google made this script which takes the average of many flags: https://plus.google.com/117808384777851490555/posts/WEBtTuDRubF
9.5. Open a web browser
import org.autoplot.AutoplotUtil org.autoplot.AutoplotUtil.openBrowser(String url)
9.6. Tweak DOM parameters not saved to VAP
Not all properties in the DOM are saved to the VAP file (anything under "controller" is not saved). As a work-around, load a VAP file and then modify the non-saved DOM elements with a script.
ds= 'file:/tmp/webtest.vap' plot( ds ) dom= getDocumentModel() dom.plotElements[0].controller.renderer.cadenceCheck = False
Save the above as /tmp/webtest.jy and enter
script:file:/tmp/webtest.jy
in the address bar. (Autoplot 2011 now has cadenceCheck in the DOM, so editing Plot Element Properties can accomplish this as well.)
Note that vaps can be embedded within a .jy script as well.
9.7. Plot to specific subpanels
Subplots may be selected:
setPlotLayout(2,3) plot(0,ripples(20)) plot(1,ripples(30)) plot(2,ripples(40))
This will add plots until the there's a spot "2" for the data. The command setPlotLayout(2,3) will make two rows of three plots each.
9.8. list remote website files
In the script panel (enabled via options):
for i in listDirectory('http://autoplot.org/data/*.cdf'): print i
9.9. make a plot for each file
In the script panel (enabled via options), enter
for i in listDirectory('http://autoplot.org/data/*.qds'): plot( 'http://autoplot.org/data/'+i ) writeToPng( '/tmp/' + i+'.png' )
9.10. Python/Jython is great with strings
product= 'c1' date= '20010101' plot( 'http://www.autoplot.org/data/%s_%s.dat' % ( product, date ) )
%s plugs in the corresponding string, %d an integer, %f a float. Often you need the result to have a fixed number of characters, so you might say %9f so that 9 characters are always used. Prefixing the number with a zero will zero-pad the field: %09d. Last the number of decimal places can be specified with: %9.2f.
print '%s' % 'string' # 'string' print '%09d' % 2 # '000000002' print '%9.3f' % 3.14 # ' 3.140'
9.11. mash data
Jython (Python in Java) scripts can be used to mash data. The getParam(parmName,default) function allows data to be passed into a script via the URI:
vap+jyds:file:///home/jbf/inbox/larry.20100212.icee/icee.jyds?type='e_star'
Here's the script:
http://autoplot.org/data/jyds/icee.jyds
Note the script can be read directly from the http web site, this one has local file references. Note too that the menu item [menubar]->Tools->Data Mash Up... allows you to mash data without code.
9.12. compare calibrated and uncalibrated data
I had data in two files that I needed to compare, but a simple calibration had been applied to one but not the other. I could easily apply the calibration with a jython script in autoplot. Instead of plotting the data, I (enable and) flip over to script tab in Autoplot and enter the script:
# code to apply calibration to compare my cdf to Dan's ds= getDataSet('vap:file:///home/jbf/temp/rbsp/fm1/L1/2041/01/14/rbsp-a_ws_emfisis-L1_20410114_v1.1.3.cdf?BuBu') ds= sqrt(ds) ds= 20 * log10( 2 * ds / 32768. ) data= ds
These scripts work by plotting the variable "data" by default. Note I can right click in the script tab and select "getDataSet" and the getDataSet command is entered with the current URI. Select "data source context" and hit the execute button, and I need to save this to a file with a .jyds extension before Autoplot can use it. I'd be able to plot this against the other data, except it has units and this new dataset doesn't. I can remove units from Dan's the same way:
# code to remove units from Dan's CDFs ds= getDataSet('vap:file:///home/jbf/temp/compareFeb3/rbsp-a_wf_emfisis-L1_20110114134618_v1.1.99.cdf?BuBu') ds= putProperty(ds,QDataSet.UNITS,None) data= ds
Now I can plot one against the other, and plot slices on the same plot. Note Autoplot will plot data of different units on the same axis, but with a warning message.
9.13. generating time ranges in Jython
In the application script dialog (Options->Enable Feature->Script Panel), you can create a list of timeranges:
trs= generateTimeRanges( '$Y-$m-$d', '2012' ) for i in trs: print i
You'll see the output in the java stdout or on the console (Options->Enable Feature->Log Console) if it's enabled.
Here's a more useful script:
# plot all datasets for the year 2012 trs= generateTimeRanges( '$Y-$m-$d', '2012' ) for i in trs: ds= 'http://cdaweb.gsfc.nasa.gov/istp_public/data/ace/swepam/level_2_cdaweb/swe_k0/$Y/ac_k0_swe_$Y$m$d_v01.cdf?He_ratio&timerange=%s' % i plot( ds ) writeToPng( '/tmp/ap/%s.png' % i )
9.14. remove the fill data from a list
Remove fill data from a list with the where and valid functions:
list= dataset( [ -1e31, 37.6, 43.2, 55.1, 97.0] ) list= putProperty( list, QDataSet.FILL_VALUE, -1e31 ) r= where( valid( list ) ) list= list[r]
9.15. interpolating dataset onto another dataset's timetags
Ivar asked about a function I've always meant to implement explicitly, but I found the current implementation is fine. He has data at timetags with one cadence, and wishes to interpolate them to another dataset's timetags. Here's the script:
# syncTimeTags.jy # Show how one dataset is synched to another. Density at 5 min resolution is loaded in, # and interpolated onto a grid of flux at roughly 4min resolution. flux4min= getDataSet( 'http://cdaweb.gsfc.nasa.gov/pub/data/ace/sis/level_2_cdaweb/sis_h1/2017/ac_h1_sis_20170117_v05.cdf?flux_He' ) density5min= getDataSet( 'http://cdaweb.gsfc.nasa.gov/pub/data/ace/mag/level_2_cdaweb/mfi_k0/2017/ac_k0_mfi_20170117_v01.cdf?Magnitude') t5min= density5min.property(QDataSet.DEPEND_0) t4min= flux4min.property(QDataSet.DEPEND_0) findx= findex( t5min, t4min ) # 5min tags interpolated to 4 minute tags density4min= interpolate( density5min, findx ) plot( 0, density5min, title= 'This is the original data' ) plot( 1, t4min, density4min, title='These line up with the flux data' ) plot( 2, flux4min, title='This is the flux' )
Here's a generic "synchronize" routine that syncs up data:
flux4min= getDataSet( 'http://cdaweb.gsfc.nasa.gov/pub/data/ace/sis/level_2_cdaweb/sis_h1/2017/ac_h1_sis_20170117_v05.cdf?flux_He' ) magn5min= getDataSet( 'http://cdaweb.gsfc.nasa.gov/pub/data/ace/mag/level_2_cdaweb/mfi_k0/2017/ac_k0_mfi_20170117_v01.cdf?Magnitude') BGSE5min= getDataSet( 'http://cdaweb.gsfc.nasa.gov/pub/data/ace/mag/level_2_cdaweb/mfi_k0/2017/ac_k0_mfi_20170117_v01.cdf?BGSEc') ( magn, BGSE ) = synchronize( flux4min, [magn5min, BGSE5min] )
Keywords: synchronize synchronizing
9.16. run test script to see that everything in history is still plottable
In Jython editor:
from test.endtoend import TryHistory TryHistory.main( [] )
This attempts to load every URI in $HOME/autoplot_data/bookmarks/history.txt as a QDataSet, printing the result and load times, and then finally reporting the statistics for the run. A future version may also plot each one to a png file.
9.17. add a Jython/Python script to the Tools menu
Jython scripts can be added to the tools menu by putting them in the bookmarks file AUTOPLOT_DATA/bookmarks/tools.xml. Set the label for the script by adding a comment to the script: "# label: My Script Label". When running a script, Autoplot will show the parameters, and there is a checkbox to add it to the menu.
9.18. Share a script with others
An application-context script can be shared with others, so that editors can pass scripts to new users. For example, type in the address bar:
script:http://autoplot.org/data/tools/flashFocus.jy
and a GUI is presented asking if you'd like to run the script. A checkbox allows the script to be added to your tools menu. Here are some other useful scripts to try (omit the #comment part):
script:http://autoplot.org/data/tools/flashFocus.jy # flash the current plot Element that's selected script:http://autoplot.org/data/tools/reloadAllUris.jy # reload all loaded data (remote files in cache will not be loaded from server if the timestamp hasn't changed) script:http://autoplot.org/data/tools/testHtmlConnection.jy # test to see if we're on line. (This should be renamed testHttpConnection.jy.) script:http://autoplot.org/data/tools/toggleDayOfYear.jy # toggle the day-of-year setting
Note scripts can contain the special comment "# label:" that gives them nice labels.
Note "*.jy" is now always a script, and the "script:" prefix is no longer needed.
9.19. Reduce a long time series
Often we need to reduce a long high-resolution time series, leaving a process running overnight. This loads the URI with the time series browse capability and saves out a reduced version. This only loads for 26 days, but it can be modified to run over years (with your data).
# title: hourly averages demo shows how to take hourly averages over a long interval # label: hourly averages from org.virbo.dsutil.Reduction import reducex import java.util.LinkedHashMap import java.lang.Exception uri= 'http://sarahandjeremy.net/~jbf/1wire/data/$Y/$m/$d/610008002FE00410.$Y$m$d.d2s?timerange=%s' dr= DatumRangeUtil.parseTimeRange('2012-09-01') endt= DatumRangeUtil.parseTimeRange('2012-09-26' ) targetRes= dataset('1 hr') # '1 min' '2 hr' '3 days'. Use Length mouse action for examples exceptions= java.util.LinkedHashMap() monitor.started() dsall= None while ( dr.min().lt( endt.max() ) ): monitor.progressMessage= "%s until %s" % ( dr, endt ) uri1= uri % dr.toString() try: ds= getDataSet( uri1 ) ds= reducex( ds, targetRes ) dsall= concatenate( dsall, ds ) # TODO: this probably won't scale, replace with dataset builder. except java.lang.Exception,ex: exceptions.put( uri1, ex ) dr= dr.next() formatDataSet( dsall, '/tmp/reduced1Hr.qds' ) print 'Exceptions encountered:' for ex in exceptions.entrySet(): print '== %s ==' % ex.getKey() print ex.getValue()
9.20. Import set of common functions
You can import a set of common functions using getFile and "execfile":
ff= getFile( 'http://emfisis.physics.uiowa.edu/pub/jy/dev/jbf/operators/experimentalFunctions.jy',monitor.getSubtaskMonitor('import')) execfile( ff.toString() ) flux4min= getDataSet( 'http://cdaweb.gsfc.nasa.gov/pub/data/ace/sis/level_2_cdaweb/sis_h1/2017/ac_h1_sis_20170117_v05.cdf?flux_He&slice1=0' ) density5min= getDataSet( 'http://cdaweb.gsfc.nasa.gov/pub/data/ace/mag/level_2_cdaweb/mfi_k0/2017/ac_k0_mfi_20170117_v01.cdf?Magnitude') dst60min= getDataSet( 'http://cdaweb.gsfc.nasa.gov/pub/data/omni/omni_cdaweb/hourly/2017/omni2_h0_mrg1hr_20170101_v01.cdf?DST') t5min= density5min.property(QDataSet.DEPEND_0) ( density, flux, dst ) = synchronize( t5min, density5min, flux4min, dst60min, nn=1 ) # experimental function (note there is a built-in function in Autoplot v2017a. plot( 0, density ) plot( 1, flux ) plot( 2, dst )
This introduces some possible security concerns, and this may be restricted in the future.
9.21. Linear Fit Routine
There's a code to does linear fits:
from org.das2.qds.util import LinFit x= linspace(0,5,40) y= x * 6 + 6 * randn(40) setLayoutOverplot(2) plot( 0, x, y ) lf= LinFit( x,y, 6*ones(40) ) plot( 1, x, x*lf.b + lf.a ) print lf.chi2 / (x.length()-1)
9.22. Arbitrary Layout
Autoplot v2015a and more recent versions allow scripts to plot to arbitrary locations on the page, bypassing the automatic layout. For example:
plot(0,rand(2000),rand(2000),xpos='20%,90%',ypos='20%,50%') plot(1,linspace(0,1,20),randn(20),xpos='50%,70%',ypos='60%,80%')
Em offsets (when an em is the current font size) can be used as well, and pixel locations:
plot(0,rand(2000),rand(2000),xpos='4em,100%-4em',ypos='4em,100%-4em') plot(1,rand(2000),rand(2000),xpos='100px,200px',ypos='100px,200px')
9.23. Add to Jython Search Path
I want to add the ability to create a video. I can add this to the search path in Jython like so:
import sys addToSearchPath(sys.path,'http://central.maven.org/maven2/org/jcodec/jcodec-javase/0.2.2/jcodec-javase-0.2.2.jar',monitor.getSubtaskMonitor('jar1')) addToSearchPath(sys.path,'http://central.maven.org/maven2/org/jcodec/jcodec/0.2.2/jcodec-0.2.2.jar',monitor.getSubtaskMonitor('jar2')) from org.jcodec.api.awt import AWTSequenceEncoder from javax.imageio import ImageIO from java.io import File enc = AWTSequenceEncoder.createSequenceEncoder(File("/home/jbf/tmp/filename.mp4"),24) dd= '/tmp/ap/kris/' ff= listDirectory(dd+'demoSound_*.png') monitor.setTaskSize(len(ff)) monitor.started() monitor.setProgressMessage('encoding the movie') for f in ff[0:10]: monitor.setTaskProgress(monitor.getTaskProgress()+1) img= ImageIO.read( File(dd+f) ) enc.encodeImage(img) enc.finish() monitor.finished()
See https://sourceforge.net/p/autoplot/feature-requests/584/.
10. Use Screenshots Tool and Pngwalk Verify
I used the pngwalk verify tool to select frames from the screenshots tool. First, I recorded my movie with the screenshots tool. In Autoplot, run http://autoplot.org/data/tools/startScreenShots.jy. This starts dumping screenshots into /home/jbf/temp/ap. I quit Autoplot when I was finished with the sequence, and then started up pngwalk and turn on the quality control. I could go through the screenshots and mark the frames I wanted to keep, which would then put .ok files next to my images. Then the unix command to extract my files was for i in *.png.ok; do echo $i; cp ${i%.ok} movie20130208; done
10.1. Bind Two PNGWalks Together
I needed to compare two pngwalks, created in two different directories but with files of the same name. A while ago I added properties to the pngwalk tool so that they could be bound together:
1. start the two pngwalk tools 2. on the console, get references to the two pngwalk tools via the app manager: * from org.autoplot import AppManager * pw1= AppManager.getInstance().getApplication(1) * pw2= AppManager.getInstance().getApplication(2) * print pw1,pw2 # to verify we got the correct references 3. and bind their "SELECTED_NAME" property: * bind( pw1, pw1.PROP_SELECTED_NAME, pw2, pw1.PROP_SELECTED_NAME )
The two pngwalks can be used to compare images now, advancing in one will advance on the other. Note the timerange property can be bound as well, for comparing pngwalks with different filenames or comparing a pngwalk to Autoplot's dom.timerange.
10.2. Make a tutorial web page
You can make effective tutorial html pages (like http://sarahandjeremy.net/~jbf/autoplot/tutorials/findJava8/20160204_1502.html) using the script: http://autoplot.org/data/tools/makeTutorialHtml.jy. This will create an html page showing slides marked as "Okay" with the QC tool. This demo shows how this is done: http://autoplot.org/autoplot/data/tutorials/20160116meta/20160116_1426.html
11. Writing to PDF files for embedding in a science poster presentation
The pixels on the desktop correspond to points when printing to PDF. So if you want a 5in by 5in figure, you should have a 360 by 360 pixel canvas (72 dpi). To get this, see the style tab, then under Canvas, Canvas Size:
- uncheck "Adjust to Fit into Application"
- Set the width to 360 (points)
- Set the height to 360 (points)
- [menubar]->Auto->"AutoLayout" can be turned off to get more control of the layout. This will disable the code that makes room for color bars and new plots.
- Plot edges can be tweaked to by holding the shift button while mousing over plot boundaries. Control blocks will appear to resize the plot.
- axes tab, Y Axis, Isotropic will lock the data:pixel ratio for the x and y axes, when the units are compatible.
12. Connecting to SAMP Hub
ESA (European Space Agency) software often uses the "SAMP Hub" to communicate data references between software packages. Autoplot has some support for the SAMP hub, being able to start a hub and accept requests. To start Autoplot with SAMP support, run Autoplot with the --samp switch turned on, like so:
/usr/local/java1.8/bin/java -jar autoplot.jar --samp
or run the script "script:http://autoplot.org/data/tools/startSampHub.jy" on the Autoplot address bar. Note this can be added to the tools menu using the checkbox at the bottom of the "Run Script" dialog.
Autoplot now includes Tools->"Start SAMP Hub" (a bookmarked tool) on new installations.
13. Analysis on each image of mp4 file
Presently Autoplot can only access png and jpg images, so you need to use ffmpeg to split up the mp4 video into jpg files:
ffmpeg -i chargeMac.mp4 /tmp/ap/frames%05d.jpg
Now Autoplot can load each (or the aggregation) using /tmp/ap/frames%x.jpg.
14. VAP files
14.1. Embedding data within a .vap file
Autoplot's save-to-vap option has a checkbox "embed data" which when checked will write a .vap.zip file, which is a zip file containing data as well as a .vap file. Any data which is local to the machine will be inserted into the zip file, and the references in the .vap will contain the macro %{PWD} which refers to the .vap file location.